Filter Image Using Frangi
SUMMARY
Filter Image Using Frangi applies Frangi vesselness filter to enhance tubular structures.
Frangi filter is designed to detect vessel-like structures in medical images, fingerprints, and other images with elongated features. This multi-scale filter uses eigenvalue analysis of the Hessian matrix to identify tubular structures while suppressing plate-like and blob-like structures. It's particularly effective in retinal imaging, angiography, and neuroimaging applications.
Use this Skill when you want to enhance and detect vessel-like tubular structures in biomedical images.
The Skill
from telekinesis import pupil
vessels = pupil.filter_image_using_frangi(
image=image,
scale_start=1,
scale_end=10,
scale_step=2,
alpha=0.5,
beta=0.5,
gamma=None,
detect_black_ridges=True,
border_type="reflect",
constant_value=0.0,
)Example
Input Image
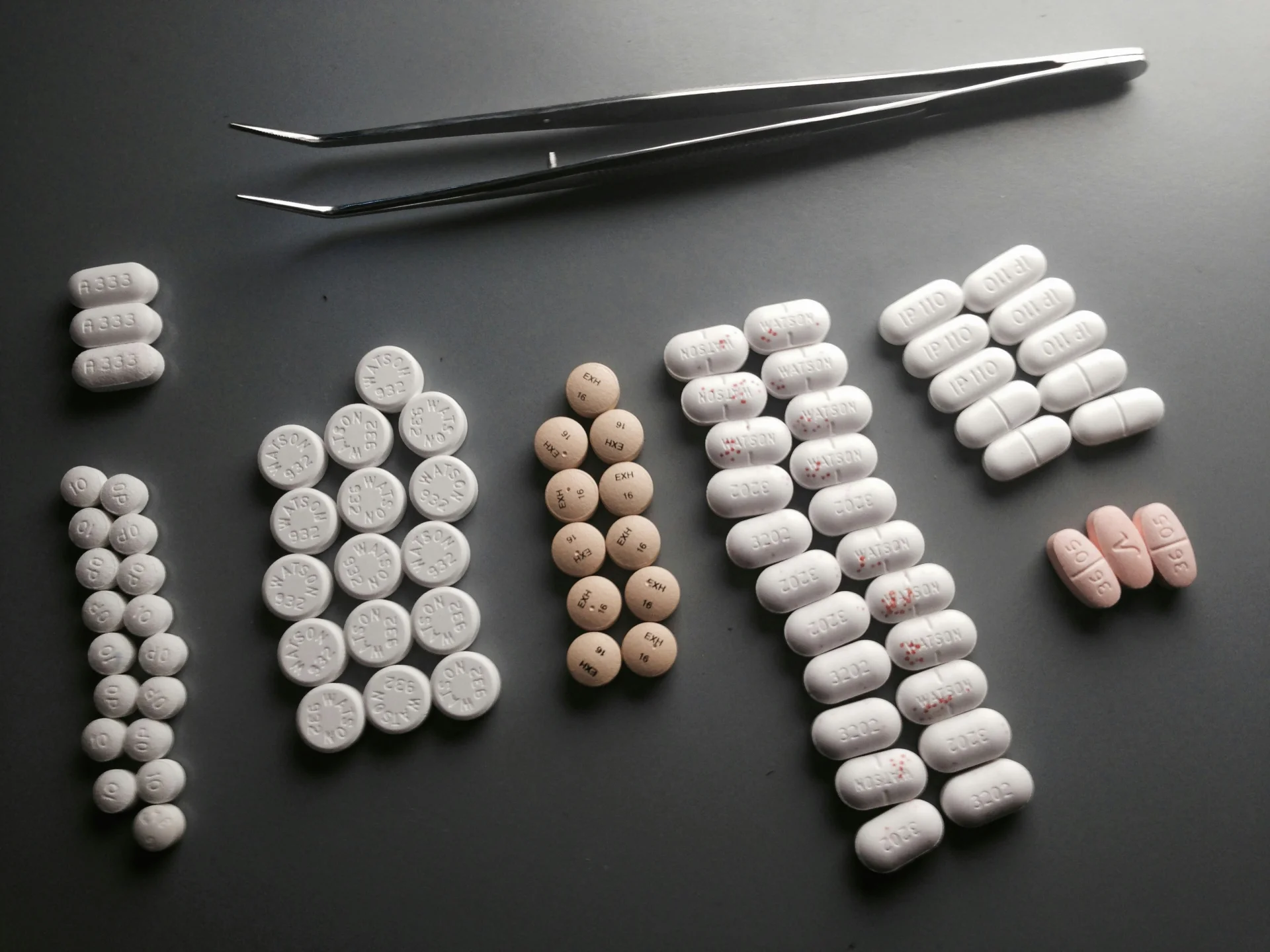
Original grayscale image with vessel structures (tablets_arranged.jpg, normalized to 0-1 range)
Filtered Image
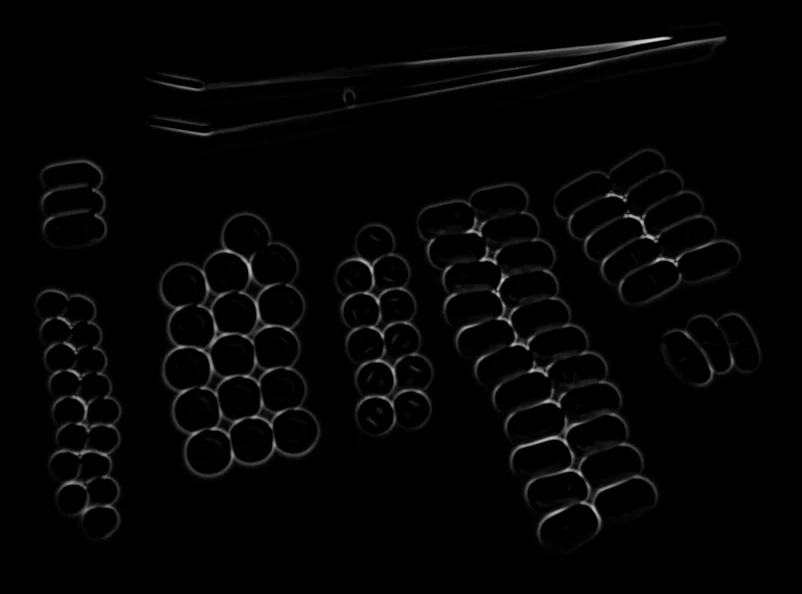

Vesselness map highlighting tubular structures
The Code
from telekinesis import pupil
from datatypes import io
import pathlib
from loguru import logger
DATA_DIR = pathlib.Path("path/to/telekinesis-data")
# Load image
filepath = str(DATA_DIR / "images" / "tablets_arranged.jpg")
image = io.load_image(filepath=filepath, as_gray=True)
logger.success(f"Loaded image from {filepath}")
# Apply Frangi filter for vesselness enhancement
filtered_image = pupil.filter_image_using_frangi(
image=image,
scale_start=6,
scale_end=10,
scale_step=1,
alpha=0.5,
beta=0.5,
detect_black_ridges=True,
border_type="reflect",
border_value=0.0,
)
# Access results
filtered_image_np = filtered_image.to_numpy()
logger.success("Applied Frangi filter. Filtered output image shape: {}", filtered_image_np.shape)The Explanation of the Code
The code begins by importing the necessary modules: pupil for image processing operations, io for data handling, pathlib for path management, and loguru for logging.
from telekinesis import pupil
from datatypes import io
import pathlib
from loguru import loggerNext, an image is loaded from a .jpg file using the io.load_image function. The image is loaded in grayscale mode using as_gray=True, which is appropriate for the Frangi filter that enhances tubular structures.
DATA_DIR = pathlib.Path("path/to/telekinesis-data")
# Load image
filepath = str(DATA_DIR / "images" / "tablets_arranged.jpg")
image = io.load_image(filepath=filepath, as_gray=True)The main operation uses the filter_image_using_frangi Skill from the pupil module. This Skill applies the Frangi vesselness filter to enhance tubular structures using multi-scale eigenvalue analysis of the Hessian matrix. The parameters can be tuned to control scale range, vessel sensitivity, and border handling depending on the characteristics of the input image.
filtered_image = pupil.filter_image_using_frangi(
image=image,
scale_start=6,
scale_end=10,
scale_step=1,
alpha=0.5,
beta=0.5,
detect_black_ridges=True,
border_type="reflect",
border_value=0.0,
)Finally, the filtered image is converted to a NumPy array using to_numpy() for further processing, visualization, or downstream tasks.
filtered_image_np = filtered_image.to_numpy()
logger.success(f"Output image shape: {filtered_image_np.shape}")This operation is particularly useful in robotics and vision pipelines for vessel detection, medical imaging, retinal imaging, and angiography, where enhancing tubular structures is required.
The output is a vesselness map, where higher values correspond to stronger tubular structure responses. The original image remains unchanged.
Running the Example
Runnable examples are available in the Telekinesis examples repository. Follow the README in that repository to set up the environment. Once set up, you can run this specific example with:
cd telekinesis-examples
python examples/pupil_examples.py --example filter_image_using_frangiHow to Tune the Parameters
The filter_image_using_frangi Skill has 9 parameters:
scale_start (default: 1):
- The minimum scale (sigma) for structure detection
- Determines the smallest vessel width to detect
- Units: Pixels (sigma)
- Increase to detect larger vessels; Decrease for finer vessels
- Typical range: 1–5 (min: 1, max: 5)
- Use 1–3 for thin vessels, 3–5 for medium vessels
scale_end (default: 10):
- The maximum scale (sigma) for structure detection
- Determines the largest vessel width to detect
- Units: Pixels (sigma)
- Increase to detect thicker vessels; Decrease for thinner range
- Typical range: 5–20 (min: 5, max: 20)
- Use 5–10 for capillaries, 10–20 for thick vessels
scale_step (default: 2):
- The step size between scales
- Smaller steps provide finer scale resolution but are slower
- Units: Pixels
- Increase for faster processing; Decrease for finer scale resolution
- Typical range: 1–5 (min: 1, max: 5)
- Use 1 for maximum precision, 2–5 for efficiency
alpha (default: 0.5):
- The weight for the blobness measure
- Increase to emphasize blob-like structures; Decrease for tubular selectivity
- Typical range: 0.1–2.0 (min: 0.1, max: 2.0)
- Use 0.1–0.5 for strict tubular detection, 0.5–2.0 for blob inclusion
beta (default: 0.5):
- The weight for the second-order structureness
- Increase to emphasize tubular structures; Decrease for less sensitivity
- Typical range: 0.1–2.0 (min: 0.1, max: 2.0)
- Use 0.5–2.0 for strong vessel enhancement
gamma (default: None):
- The weight for background suppression. If None, computed automatically
- Typical range: 0.1–2.0 when set explicitly
- Use None for automatic; set explicitly to tune background sensitivity
detect_black_ridges (default: True):
- Whether to detect dark ridges (vessels) instead of bright
- Options: True, False
- Use True for dark vessels on bright background (medical imaging); False for bright on dark
border_type (default: "reflect"):
- The padding mode for ridge detection
- Options: "reflect", "constant", "edge", "symmetric", "wrap"
- Use "reflect" for most cases; "constant" with border_value for custom padding
border_value (default: 0.0):
- The value used for constant padding mode
- Units: Intensity (0–1 range)
- Typical range: 0.0–1.0 (min: 0.0, max: 1.0)
- Use 0.0 for black padding when border_type="constant"
TIP
Best practice: Set scale_start and scale_end to span the expected range of vessel widths in your image. Use scale_step=2 for efficiency, or 1 for maximum precision.
Where to Use the Skill
Filter Using Frangi is commonly used in the following pipelines:
- Retinal vessel segmentation - Detect blood vessels in retinal images
- Angiography analysis - Enhance vascular structures in medical scans
- Fingerprint enhancement - Enhance ridge structures
- Neuroimaging - Detect neural fibers and blood vessels
- Quality control - Detect cracks, scratches, or linear defects
Related skills to build such a pipeline:
filter_image_morphological_open: Clean up vessel detection resultsfilter_image_using_hessian: Alternative vesselness measure
Alternative Skills
| Skill | vs. Filter Using Frangi |
|---|---|
| filter_image_using_hessian | Hessian vesselness is simpler. Use Frangi for more sophisticated vessel detection, Hessian for speed. |
| filter_image_using_sato | Sato is optimized for ridge detection. Use Frangi for vessels, Sato for general ridges. |
| filter_image_using_meijering | Meijering is optimized for neurites. Use Frangi for general vessels, Meijering for fine branching structures. |
When Not to Use the Skill
Do not use Filter Using Frangi when:
- You need simple edge detection (Use Sobel or Canny instead)
- You're detecting non-tubular structures (Use appropriate edge or blob detectors)
- You need very fast processing (Frangi is computationally expensive due to multi-scale analysis)
- Your structures are not vessel-like (Use general-purpose filters instead)

